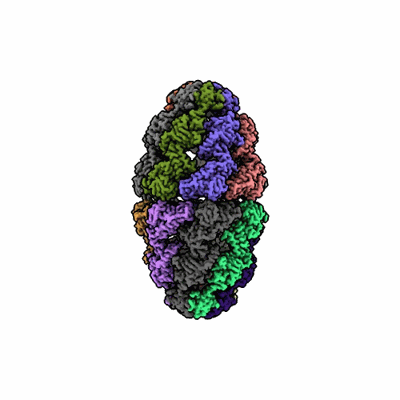
Cryo-EM structure of H. thermoluteolus GroEL-GroES2 football complex
Resolution: 2.5 Å
EM Method: Single-particle
Fitted PDBs: 8wuc
Q-score: 0.554
Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S
Structure (2024) [ DOI: doi:10.1016/j.str.2024.02.012 Pubmed: 38492570 ]
- H. thermoluteolus groel (Complex)
- Chaperonin groel (56 kDa, Protein from Hydrogenophilus thermoluteolus)
- H. thermoluteolus groes (Complex)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Co-chaperonin groes (10 kDa, Protein from Hydrogenophilus thermoluteolus)
- Magnesium ion (24 Da, Ligand)
- H. thermoluteolus groel-groes2 football complex bound with adp-mg2+ (73 MDa, Complex from Hydrogenophilus thermoluteolus)
Cryo-EM structure of H. thermophilus GroEL-GroES2 asymmetric football complex
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8wuw
Q-score: 0.444
Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S
Structure (2024) [ DOI: doi:10.1016/j.str.2024.02.012 Pubmed: 38492570 ]
- Potassium ion (39 Da, Ligand)
- H. thermophilus groel-groes2 asymmetric football complex (Complex from Hydrogenobacter thermophilus TK-6)
- Chaperonin groel (Complex)
- Co-chaperonin groes (Complex)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Co-chaperonin groes (10 kDa, Protein from Hydrogenobacter thermophilus TK-6)
- Chaperonin groel (56 kDa, Protein from Hydrogenobacter thermophilus TK-6)
Cryo-EM structure of H. thermophilus GroEL-GroES bullet complex
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8wux
Q-score: 0.453
Liao Z, Gopalasingam CC, Kameya M, Gerle C, Shigematsu H, Ishii M, Arakawa T, Fushinobu S
Structure (2024) [ DOI: doi:10.1016/j.str.2024.02.012 Pubmed: 38492570 ]
- Co-chaperonin groes (10 kDa, Protein from Hydrogenobacter thermophilus TK-6)
- Potassium ion (39 Da, Ligand)
- H. thermophilus groel-groes bullet complex (Complex from Hydrogenobacter thermophilus TK-6)
- Chaperonin groel (Complex)
- Phosphoaminophosphonic acid-adenylate ester (506 Da, Ligand)
- Magnesium ion (24 Da, Ligand)
- Co-chaperonin groes (Complex)
- Chaperonin groel (56 kDa, Protein from Hydrogenobacter thermophilus TK-6)
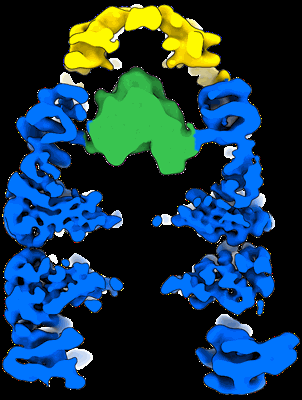
CryoEM structure of GroEL-GroES-ADP.AlF3-Rubisco.
Resolution: 3.7 Å
EM Method: Single-particle
Fitted PDBs: 8ba9
Q-score: 0.395
Gardner S, Darrow MC, Lukoyanova N, Thalassinos K, Saibil HR
PNAS (2023) 120 pp. e2308933120- [ DOI: doi:10.1073/pnas.2308933120 Pubmed: 38064510 ]
- Potassium ion (39 Da, Ligand)
- Aluminum fluoride (83 Da, Ligand)
- Co-chaperonin groes (10 kDa, Protein from Escherichia coli K-12)
- 60 kda chaperonin (55 kDa, Protein from Escherichia coli K-12)
- Adenosine-5'-diphosphate (427 Da, Ligand)
- Groel-adp.befx-rubisco (802 kDa, Complex from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
CryoEM reconstruction of GroEL-GroES-ADP.AlF3-Rubisco, class I.
Gardner S, Darrow MC, Lukoyanova N, Thalassinos K, Saibil HR
PNAS (2023) 120 pp. e2308933120-e2308933120 [ DOI: doi:10.1073/pnas.2308933120 Pubmed: 38064510 ]
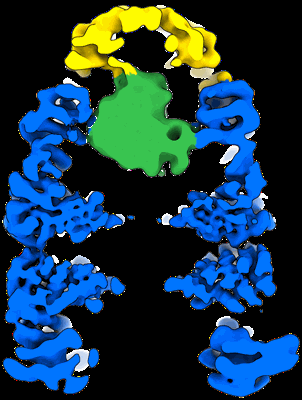
CryoEM reconstruction of GroEL-GroES-ADP.AlF3-Rubisco, class III.
Gardner S, Darrow MC, Lukoyanova N, Thalassinos K, Saibil HR
PNAS (2023) 120 pp. e2308933120-e2308933120 [ DOI: doi:10.1073/pnas.2308933120 Pubmed: 38064510 ]
- Groel-groes-adp.alf3-rubisco (802 kDa, Complex from Escherichia coli)
CryoEM reconstruction of GroEL-GroES-ADP.AlF3-Rubisco, class IV.
Gardner S, Darrow MC, Lukoyanova N, Thalassinos K, Saibil HR
PNAS (2023) 120 pp. e2308933120-e2308933120 [ DOI: doi:10.1073/pnas.2308933120 Pubmed: 38064510 ]
- Groel-groes-adp.alf3-rubisco (802 kDa, Complex from Escherichia coli)
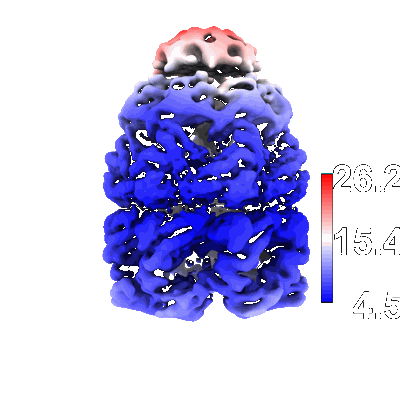
Map of GroEL:GroES-(ADP) complex plunge frozen 200 ms after reaction initiation with ATP
Torino S, Dhurandhar M, Stroobants A, Claessens R, Efremov RG
Nat Methods (2023) 20 pp. 1400-1408 [ Pubmed: 37592181 DOI: doi:10.1038/s41592-023-01967-z ]
- Groes (Protein from Escherichia coli)
- Groel (Protein from Escherichia coli)
- Hetero 14mer assembled from 2 heptameric rings (950 kDa, Complex from Escherichia coli)
Movies of GroEL-ES-nucleotide complex plunged 13, 50, 200 ms and 20 s after mixing with ATP
Torino S, Dhurandhar M, Stroobants A, Claessens R, Efremov RG
Nat Methods () [ DOI: 10.1038/s41592-023-01967-z ]
Image sets:
- GroEL 13ms timepoint
- GroELES 200ms timepoint
- GroELES with 20s of mixing time
- GroELES at 50ms timepoint
GroEL:GroES-ATP complex under continuous turnover condirions
Resolution: 2.3 Å
EM Method: Single-particle
Fitted PDBs: 8bkz
Q-score: 0.632
Torino S, Dhurandhar M, Stroobants A, Claessens R, Efremov RG
Nat Methods (2023) 20 pp. 1400-1408 [ Pubmed: 37592181 DOI: doi:10.1038/s41592-023-01967-z ]
- Co-chaperonin groes (10 kDa, Protein from Escherichia coli)
- Potassium ion (39 Da, Ligand)
- Hetero 14mer assembled from 2 heptameric rings (950 kDa, Complex from Escherichia coli)
- Water (18 Da, Ligand)
- Adenosine-5'-triphosphate (507 Da, Ligand)
- Chaperonin groel (57 kDa, Protein from Escherichia coli)
- Magnesium ion (24 Da, Ligand)