Filter By
Current Database
Current status
Sample type
Organism
- Escherichia coli [841]
- Homo sapiens [741]
- Saccharomyces cerevisiae [431]
- Escherichia coli [257]
- Oryctolagus cuniculus [198]
- Saccharomyces cerevisiae [159]
- Synthetic construct [154]
- Mus musculus [80]
- Chaetomium thermophilum [77]
- Bacillus subtilis [53]
- Mycoplasma pneumoniae [52]
- Escherichia coli [51]
- Bos taurus [42]
- Canis lupus [40]
- Thermus thermophilus [34]
- Schizosaccharomyces pombe [33]
- Mycolicibacterium smegmatis [30]
- Saccharomyces cerevisiae [29]
- Sus scrofa [26]
- Bacillus subtilis [24]
- Listeria innocua [24]
- Staphylococcus aureus [23]
- Triticum aestivum [22]
- Spinacia oleracea [20]
- Acinetobacter baumannii [20]
- Taura syndrome virus [19]
- Thermus thermophilus [19]
- Severe acute respiratory syndrome coronavirus [18]
- Staphylococcus aureus [18]
- Rattus norvegicus [17]
- Trypanosoma brucei [17]
- Saccharomyces cerevisiae [17]
- Cricket paralysis virus [16]
- Enterococcus faecalis [14]
- Kluyveromyces lactis [14]
- Dictyostelium discoideum [14]
- (no species) [13]
- Hepacivirus c [13]
- Mycobacterium tuberculosis [13]
- Sulfolobus acidocaldarius [13]
- Chaetomium thermophilum [12]
- Caenorhabditis elegans [12]
- Xenopus laevis [12]
- Hepacivirus c [11]
- Mycolicibacterium smegmatis [11]
- Chlamydomonas reinhardtii [11]
- Tetrahymena thermophila [11]
- Drosophila melanogaster [11]
- Mycobacterium tuberculosis [11]
- Pseudomonas aeruginosa [10]
- Enterobacteria phage t4 [10]
- Dictyostelium discoideum [9]
- Acinetobacter baumannii [9]
- Plasmodium falciparum [9]
- Listeria monocytogenes [8]
- Giardia intestinalis [8]
- Vairimorpha necatrix [8]
- Apis mellifera [8]
- Bacteroides thetaiotaomicron [7]
- Kluyveromyces lactis [7]
- Pseudomonas aeruginosa [7]
- Solanum lycopersicum [7]
- Thermomyces lanuginosus [7]
- Pyrococcus abyssi [6]
- Israeli acute paralysis virus [6]
- Triticum sp. [6]
- Escherichia coli [6]
- Giardia intestinalis [6]
- Sus scrofa [6]
- Avian coronavirus [5]
- Mycoplasma pneumoniae [5]
- Candida albicans [5]
- Escherichia coli [5]
- Escherichia coli [5]
- Trypanosoma cruzi [5]
- Acinetobacter baumannii [5]
- Escherichia coli [5]
- Acinetobacter sp. cip 102082 [4]
- Acinetobacter sp. cip 51.11 [4]
- Acinetobacter beijerinckii [4]
- Acinetobacter sp. niph 899 [4]
- Escherichia coli [4]
- Acinetobacter sp. 809848 [4]
- Acinetobacter sp. 263903-1 [4]
- Streptococcus pneumoniae [4]
- Cutibacterium acnes [4]
- Acinetobacter sp. anc 5600 [4]
- Acinetobacter sp. anc 4470 [4]
- Thermus thermophilus [4]
- Escherichia phage lambda [4]
- Thermococcus kodakarensis [4]
- Hepacivirus c [4]
- Neurospora crassa [4]
- Bacteriophage sp. [4]
- Acinetobacter rudis [4]
- Oryza sativa [4]
- Enterococcus faecalis [4]
- Neurospora crassa [4]
- Acinetobacter venetianus [4]
- Candida albicans [4]
Resolution
Model molecular weight
Release Date
Author by name
- Beckmann R [382]
- Berninghausen O [305]
- Williamson JR [190]
- Frank J [181]
- Cheng J [178]
- Lyumkis D [169]
- Mielke T [160]
- Wilson DN [154]
- Klinge S [150]
- Becker T [137]
- Vanden Broeck A [137]
- Amunts A [131]
- Hurt E [127]
- Spahn CMT [126]
- Thoms M [124]
- Rabuck-gibbons JN [121]
- Burger J [109]
- Ban N [106]
- Korostelev AA [105]
- Ramakrishnan V [105]
- Dong X [101]
- Li N [101]
- Sheng K [101]
- Mahamid J [96]
- Lau B [95]
- Rodnina MV [90]
- Cramer P [89]
- Gao N [89]
- Itoh Y [78]
- Forster F [77]
- Kellner N [75]
- Pfeffer S [74]
- Beck M [73]
- Demo G [72]
- Flemming D [70]
- Buschauer R [69]
- Denk T [68]
- Boehringer D [67]
- Xue L [67]
- Loveland AB [66]
- Myasnikov AG [63]
- Fischer P [62]
- Fraser JS [62]
- Lee DJ [61]
- Brown ZP [60]
- Brown A [59]
- Leibundgut M [59]
- Falquet L [57]
- Fromm L [57]
- Hashem Y [57]
- Tegunov D [57]
- Pellegrino J [56]
- Schacherl M [56]
- Sprink T [56]
- Pestova TV [55]
- Seiple IB [55]
- Inada T [54]
- Li X [54]
- Hellen CUT [53]
- Beckert B [52]
- Dabrowski M [52]
- Fedry J [52]
- Gagnon MG [52]
- Khusainov I [52]
- Loerke J [52]
- Ameismeier M [51]
- Matsuo Y [51]
- Baumeister W [50]
- Flugel T [49]
- Grigorieff N [49]
- Chen Y [48]
- Ehrenberg M [47]
- Wang C [47]
- Yusupov M [47]
- Abaeva IS [46]
- Crowe-mcauliffe C [46]
- Hegde RS [46]
- Li W [45]
- Unbehaun A [45]
- Bock LV [44]
- De S [44]
- Fernandez IS [44]
- Fluegel T [44]
- Green R [44]
- Ortega J [44]
- Rappsilber J [44]
- Zhang Y [44]
- Fischer N [43]
- Schroeer B [43]
- Diebolder CA [42]
- Guillen Schlippe Yv [42]
- Hauryliuk V [42]
- Jomaa A [42]
- Lenz S [42]
- Zhang J [42]
- Fucini P [41]
- Kreysing JP [41]
- Welsch S [41]
- Cate JHD [40]
- Carragher B [39]
Author by ORCID
- Beckmann R [194]
- Williamson JR [158]
- Klinge S [145]
- Vanden Broeck A [135]
- Cheng J [126]
- Wilson DN [122]
- Lyumkis D [121]
- Korostelev AA [103]
- Hurt E [95]
- Lau B [95]
- Ramakrishnan V [85]
- Amunts A [84]
- Gao N [84]
- Thoms M [83]
- Mahamid J [82]
- Beck M [73]
- Demo G [72]
- Berninghausen O [70]
- Flemming D [69]
- Denk T [68]
- Cramer P [65]
- Li N [64]
- Pfeffer S [63]
- Frank J [61]
- Burger J [61]
- Mielke T [57]
- Fromm L [57]
- Xue L [57]
- Falquet L [56]
- Boehringer D [53]
- Brown A [52]
- Spahn CMT [52]
- Becker T [50]
- Matsuo Y [50]
- Brown ZP [50]
- Khusainov I [48]
- Forster F [46]
- Loerke J [45]
- Hellen CUT [44]
- Pestova TV [44]
- Rappsilber J [43]
- Rabuck-gibbons JN [43]
- Rodnina MV [43]
- Lenz S [42]
- Kreysing JP [41]
- Welsch S [41]
- Grigorieff N [40]
- Fernandez IS [39]
- Gagnon MG [39]
- Hegde RS [38]
- Yokoyama T [38]
- Zimmermann-kogadeeva M [36]
- Bork P [36]
- Des Georges A [36]
- Diebolder CA [35]
- Guillen Schlippe Yv [35]
- Fraser JS [35]
- Flugel T [35]
- Loveland AB [34]
- Scheerer P [33]
- Shao S [33]
- Cate JHD [32]
- Li N [32]
- Sun M [32]
- Buschauer R [32]
- Blanchard SC [32]
- Faller WJ [31]
- Jomaa A [31]
- Plitzko JM [31]
- Hilal T [31]
- Hauryliuk V [31]
- Ebright RH [30]
- Kaelber JT [30]
- Bhushan S [30]
- Muller CW [30]
- Bridges HR [29]
- Itoh Y [29]
- Hirst J [29]
- Wang C [29]
- Molodtsov V [29]
- Turonova B [28]
- Pellegrino J [28]
- Xing H [28]
- Seiple IB [28]
- Ikeuchi K [28]
- Crowe-mcauliffe C [28]
- Taniguchi R [28]
- Bock LV [28]
- Qin B [27]
- Watson ZL [27]
- Nikolay R [27]
- Vieira-vieira CH [26]
- Tesina P [26]
- Myasnikov AG [26]
- Blaza JN [25]
- Mao Y [25]
- Llacer JL [24]
- Hou YM [24]
- Blaha G [24]
- Valle M [24]
Journal
- NAT COMMUN [581]
- SCIENCE [389]
- NATURE [388]
- MOL CELL [250]
- TO BE PUBLISHED [195]
- NUCLEIC ACIDS RES [191]
- ELIFE [176]
- NAT STRUCT MOL BIOL [171]
- PNAS [161]
- STRUCTURE [134]
- CELL [129]
- EMBO J [121]
- BIORXIV [98]
- NAT METHODS [72]
- EMBO REP [71]
- SCI ADV [51]
- CELL REP [47]
- J MOL BIOL [26]
- J STRUCT BIOL [26]
- PLOS BIOL [25]
- PLOS ONE [25]
- RNA [24]
- SCI REP [24]
- ACTA CRYSTALLOGR D STRUCT BIOL [23]
- NAT CHEM BIOL [23]
- MBIO [19]
- J BIOL CHEM [18]
- BIOMOLECULES [17]
- NAT MICROBIOL [17]
- CELL RES [16]
- PROTEIN CELL [13]
- NAT PLANTS [12]
- INT J MOL SCI [8]
- IUCRJ [8]
- PLANT COMMUN. [7]
- ACS CHEM BIOL [6]
- ANTIOXID REDOX SIGNAL [6]
- J.NEUROSCI. [6]
- PROTEIN SCI [6]
- PROTEOMICS [6]
- ACS CENT SCI [5]
- COMMUN BIOL [5]
- MSPHERE [5]
- ACS PHARMACOL TRANSL SCI [4]
- BIOPOLYM CELL [4]
- CELL DISCOV [4]
- J AM CHEM SOC [4]
- J BACTERIOL [4]
- J CELL BIOL [4]
- J.BIOCHEM. [4]
- NAT CHEM [4]
- PNAS NEXUS [4]
- BIOCHEM J [3]
- FRONT MOL BIOSCI [3]
- ISCIENCE [3]
- RES COMPUT MOL BIOL [3]
- STRUCTURE (LONDON, ENGLAND : 1993) [3]
- ULTRAMICROSCOPY [3]
- BIOCHEMISTRY [2]
- CANCER DISCOV [2]
- CHEM MED CHEM [2]
- MICROBIOL SPECTR [2]
- NAT CELL BIOL [2]
- NAT PROTOC [2]
- PROTEINS [2]
- RNA BIOL [2]
- BIOENGINEERING (BASEL) [1]
- CYSTAL RESEARCH AND TECHNOLOGY [1]
- EMERG MICROBES INFECT [1]
- FEBS LETT [1]
- GENOME BIOL EVOL [1]
- J MED CHEM [1]
- LIFE SCI ALLIANCE [1]
- METHODS [1]
- METHODS IN MOLECULAR BIOLOGY (CLIFTON, N.J.) [1]
- METHODS MOL BIOL [1]
- MICROBIOLOGY (READING, ENGL.) [1]
- MOL CELL PROTEOMICS [1]
- MOLECULES [1]
- NANOMATERIALS (BASEL) [1]
- NUC. ACIDS RES. [1]
- NUCLEIC ACIDS RESEARCH [1]
Software
- RELION [1324]
- CRYOSPARC [491]
- CTFFIND [273]
- GCTF [217]
- SPIDER [176]
- FREALIGN [119]
- WARP [105]
- PYTOM [72]
- CHIMERA [52]
- CISTEM [46]
- MDFF [36]
- IMOD [35]
- COOT [33]
- CTFFIND3, RELION [22]
- EMAN [20]
- SPIDER, APPION [17]
- IMAGIC, CUSTOM, SPIDER [16]
- APPION, SPIDER, EMAN [15]
- O [15]
- XMIPP [14]
- CTFFIND3 [13]
- UCSF CHIMERA [13]
- SPIDER, RELION [12]
- EMAN2 [11]
- GAP [11]
- PHENIX [11]
- RELION, CTFFIND3 [11]
- SIGNATURE, CTFFIND3, SPIDER, SPARX [11]
- SPIDER PACKAGE [11]
- UNBLUR, CTFFIND4, FREALIGN [11]
- CHIMERA, COOT [10]
- IMAGIC [10]
- SPIDER, SPARX [10]
- EMAN2, IMAGIC, FREALIGN, RSAMPLE, CTFFIND3 [9]
- SITUS [9]
- EMCLARITY [8]
- SPIDER, PACKAGE [8]
- PEET [7]
- PYTOM, TOM_TOOLBOX, AV3_TOOLBOX [7]
- TOM TOOLBOX [7]
- XMIPP, SPIDER [7]
- AV3, TOM, PYTOM [6]
- SPHIRE [6]
- STOPGAP [6]
- URO [6]
- CHIMERA, CNS [5]
- CHIMERA, COOT, MDFF [5]
- CHIMERA, REFMAC [5]
- DYNAMO [5]
- IMAGIC, EMAN, FREALIGN, SPIDER, GAP [5]
- IMAGIC-5, SPIDER [5]
- SPIDER, IMAGIC, GAP, CNS, XPLOR [5]
- CHIMERA, EMFIT, COOT [4]
- EM [4]
- NAMD [4]
- RSREF2.0 [4]
- SPIDER, SIGNATURE, MAPPOS [4]
- SPIDER/WEB [4]
- XMIPP, FREALIGN [4]
- ARACHNID, CTFFIND3, RELION [3]
- ARACHNID, RELION, SPIDER [3]
- CHIMERA, COOT, REFMAC [3]
- CHIMERA, MDFF [3]
- CNS [3]
- CRYOLO [3]
- CTFFIND3, RELION1.3 [3]
- IMAGIC 5 [3]
- MANUAL [3]
- MOTIONCORR2 [3]
- SIGNATURE, SPIDER [3]
- SIMPLE [3]
- SPIDER AND GAP [3]
- SPIDER XMIPP [3]
- SPIDER, WELLMAP [3]
- TOM, AV3, PYTOM [3]
- CHIMERA, O, COOT [2]
- CRYODRGN [2]
- CTFFIND3, EMAN2.1, RELION1.3 [2]
- CTFFIND3, RELION, 1.2 [2]
- CTFFIND3, RELION, 1.2, 1.3 [2]
- CTFFIND4 [2]
- EMAN2, CTFFIND4, SPIDER, SPARX [2]
- GROMACS [2]
- IMAGIC-5 SPIDER [2]
- IMAGIC-V [2]
- ML3D [2]
- MOLECULAR DYNAMICS BASED FLEXIBLE FITTING MDFF [2]
- PROGRAM O [2]
- PYMOL, MDFF [2]
- S2S, MODERNA, MODELLER, MDFF [2]
- SITUS, LOCALFIT [2]
- SPARX [2]
- TNT [2]
- UCSF-CHIMERA, MDFF [2]
- URO, REFMAC, NMFF [2]
- AMIRA 3.1, TGS SOFTWARE [1]
- APPION [1]
- AV3 [1]
- BKPR [1]
- CCP4 PACKAGE [1]
Microscope
- FEI TITAN KRIOS [2508]
- FEI POLARA 300 [196]
- FEI TALOS ARCTICA [150]
- TFS KRIOS [146]
- FEI TECNAI F20 [138]
- FEI TECNAI F30 [101]
- FEI/PHILIPS CM200FEG [42]
- FEI TECNAI ARCTICA [26]
- FEI TECNAI 20 [20]
- TFS GLACIOS [20]
- FEI TECNAI SPIRIT [18]
- JEOL 2010F [17]
- FEI TITAN [14]
- FEI TECNAI 12 [10]
- JEOL 2200FS [9]
- JEOL 3200FS [8]
- JEOL CRYO ARM 200 [8]
- JEOL CRYO ARM 300 [8]
- TFS TALOS [6]
- FEI/PHILIPS CM300FEG/T [4]
- JEOL 3100FFC [4]
- JEOL 3200FSC [4]
- FEI/PHILIPS CM200FEG/ST [2]
- JEOL 2200FSC [2]
- JEOL KYOTO-3000SFF [2]
- OTHER [2]
- FEI/PHILIPS CM200FEG/SOPHIE [1]
- FEI/PHILIPS CM200FEG/UT [1]
- FEI/PHILIPS EM420 [1]
- JEOL 1400/HR + YPS FEG [1]
- JEOL 2100F [1]
- JEOL 4000EX [1]
Electron Source
Film or detector model
- GATAN K2 SUMMIT (4K X 4K) [1258]
- GATAN K3 (6K X 4K) [525]
- FEI FALCON II (4K X 4K) [369]
- GATAN K3 BIOQUANTUM (6K X 4K) [323]
- FEI FALCON III (4K X 4K) [272]
- KODAK SO-163 FILM [162]
- GATAN K2 QUANTUM (4K X 4K) [104]
- FEI FALCON IV (4K X 4K) [92]
- TVIPS TEMCAM-F416 (4K X 4K) [50]
- FEI EAGLE (4K X 4K) [37]
- GATAN ULTRASCAN 4000 (4K X 4K) [36]
- GATAN K2 (4K X 4K) [34]
- FEI FALCON I (4K X 4K) [25]
- GENERIC TVIPS (4K X 4K) [23]
- TVIPS TEMCAM-F415 (4K X 4K) [21]
- OTHER [20]
- GATAN K2 BASE (4K X 4K) [17]
- DIRECT ELECTRON DE-64 (8K X 8K) [12]
- DIRECT ELECTRON DE-20 (5K X 3K) [10]
- GATAN MULTISCAN [7]
- GENERIC CCD [5]
- GENERIC GATAN (4K X 4K) [5]
- TVIPS TEMCAM-F816 (8K X 8K) [5]
- DIRECT ELECTRON DE-16 (4K X 4K) [4]
- GATAN K2 IS (4K X 4K) [4]
- DIRECT ELECTRON DE-12 (4K X 3K) [3]
- FEI EAGLE (2K X 2K) [3]
- GENERIC GATAN [3]
- KODAK 4489 FILM [2]
- DECTRIS SINGLA (1K X 1K) [1]
- GATAN ULTRASCAN 1000 (2K X 2K) [1]
- GENERIC FILM [1]
- TFS FALCON 4I (4K X 4K) [1]
EMPIAR Experiment type
EMPIAR Image category
Xue L, Spahn C, Schacherl M, Mahamid J
To Be Published
Xue L, Spahn C, Schacherl M, Mahamid J
To Be Published
Xue L, Spahn C, Schacherl M, Mahamid J
To Be Published
Xue L, Spahn C, Schacherl M, Mahamid J
To Be Published
Xue L, Spahn C, Schacherl M, Mahamid J
To Be Published
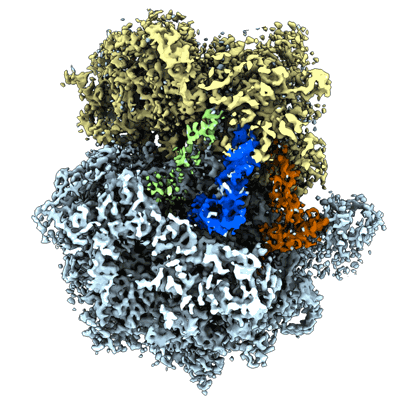
The global structure of pre50S related to DbpA in state3
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8j1z
Q-score: 0.582
Yu T, Jiang J, Yu Q, Li X, Dong W, Zeng F
To Be Published
- 5s rrna (38 kDa, RNA from Escherichia coli)
- 50s ribosomal protein l32 (6 kDa, Protein from Escherichia coli)
- Magnesium ion (24 Da, Ligand)
- 50s ribosomal protein l19 (13 kDa, Protein from Escherichia coli)
- 23s rrna (941 kDa, RNA from Escherichia coli)
- 50s ribosomal protein l23 (11 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l28 (9 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l20 (13 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l34 (5 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l18 (12 kDa, Protein from Escherichia coli)
- Water (18 Da, Ligand)
- 50s ribosomal protein l35 (7 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l22 (12 kDa, Protein from Escherichia coli)
- Ribosome 50s subunit (Complex from Escherichia coli)
- 50s ribosomal protein l27 (9 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l17 (14 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l14 (13 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l3 (22 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l33 (6 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l24 (11 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l9 (15 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l29 (7 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l13 (16 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l15 (15 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l2 (29 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l30 (6 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l5 (20 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l21 (11 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l4 (22 kDa, Protein from Escherichia coli)
- 50s ribosomal protein l25 (10 kDa, Protein from Escherichia coli)
Filter By
Current Database
Current status
Sample type
Organism
- Escherichia coli [841]
- Homo sapiens [741]
- Saccharomyces cerevisiae [431]
- Escherichia coli [257]
- Oryctolagus cuniculus [198]
- Saccharomyces cerevisiae [159]
- Synthetic construct [154]
- Mus musculus [80]
- Chaetomium thermophilum [77]
- Bacillus subtilis [53]
- Mycoplasma pneumoniae [52]
- Escherichia coli [51]
- Bos taurus [42]
- Canis lupus [40]
- Thermus thermophilus [34]
- Schizosaccharomyces pombe [33]
- Mycolicibacterium smegmatis [30]
- Saccharomyces cerevisiae [29]
- Sus scrofa [26]
- Bacillus subtilis [24]
- Listeria innocua [24]
- Staphylococcus aureus [23]
- Triticum aestivum [22]
- Spinacia oleracea [20]
- Acinetobacter baumannii [20]
- Taura syndrome virus [19]
- Thermus thermophilus [19]
- Severe acute respiratory syndrome coronavirus [18]
- Staphylococcus aureus [18]
- Rattus norvegicus [17]
- Trypanosoma brucei [17]
- Saccharomyces cerevisiae [17]
- Cricket paralysis virus [16]
- Enterococcus faecalis [14]
- Kluyveromyces lactis [14]
- Dictyostelium discoideum [14]
- (no species) [13]
- Hepacivirus c [13]
- Mycobacterium tuberculosis [13]
- Sulfolobus acidocaldarius [13]
- Chaetomium thermophilum [12]
- Caenorhabditis elegans [12]
- Xenopus laevis [12]
- Hepacivirus c [11]
- Mycolicibacterium smegmatis [11]
- Chlamydomonas reinhardtii [11]
- Tetrahymena thermophila [11]
- Drosophila melanogaster [11]
- Mycobacterium tuberculosis [11]
- Pseudomonas aeruginosa [10]
- Enterobacteria phage t4 [10]
- Dictyostelium discoideum [9]
- Acinetobacter baumannii [9]
- Plasmodium falciparum [9]
- Listeria monocytogenes [8]
- Giardia intestinalis [8]
- Vairimorpha necatrix [8]
- Apis mellifera [8]
- Bacteroides thetaiotaomicron [7]
- Kluyveromyces lactis [7]
- Pseudomonas aeruginosa [7]
- Solanum lycopersicum [7]
- Thermomyces lanuginosus [7]
- Pyrococcus abyssi [6]
- Israeli acute paralysis virus [6]
- Triticum sp. [6]
- Escherichia coli [6]
- Giardia intestinalis [6]
- Sus scrofa [6]
- Avian coronavirus [5]
- Mycoplasma pneumoniae [5]
- Candida albicans [5]
- Escherichia coli [5]
- Escherichia coli [5]
- Trypanosoma cruzi [5]
- Acinetobacter baumannii [5]
- Escherichia coli [5]
- Acinetobacter sp. cip 102082 [4]
- Acinetobacter sp. cip 51.11 [4]
- Acinetobacter beijerinckii [4]
- Acinetobacter sp. niph 899 [4]
- Escherichia coli [4]
- Acinetobacter sp. 809848 [4]
- Acinetobacter sp. 263903-1 [4]
- Streptococcus pneumoniae [4]
- Cutibacterium acnes [4]
- Acinetobacter sp. anc 5600 [4]
- Acinetobacter sp. anc 4470 [4]
- Thermus thermophilus [4]
- Escherichia phage lambda [4]
- Thermococcus kodakarensis [4]
- Hepacivirus c [4]
- Neurospora crassa [4]
- Bacteriophage sp. [4]
- Acinetobacter rudis [4]
- Oryza sativa [4]
- Enterococcus faecalis [4]
- Neurospora crassa [4]
- Acinetobacter venetianus [4]
- Candida albicans [4]
Resolution
Model molecular weight
Release Date
Author by name
- Beckmann R [382]
- Berninghausen O [305]
- Williamson JR [190]
- Frank J [181]
- Cheng J [178]
- Lyumkis D [169]
- Mielke T [160]
- Wilson DN [154]
- Klinge S [150]
- Becker T [137]
- Vanden Broeck A [137]
- Amunts A [131]
- Hurt E [127]
- Spahn CMT [126]
- Thoms M [124]
- Rabuck-gibbons JN [121]
- Burger J [109]
- Ban N [106]
- Korostelev AA [105]
- Ramakrishnan V [105]
- Dong X [101]
- Li N [101]
- Sheng K [101]
- Mahamid J [96]
- Lau B [95]
- Rodnina MV [90]
- Cramer P [89]
- Gao N [89]
- Itoh Y [78]
- Forster F [77]
- Kellner N [75]
- Pfeffer S [74]
- Beck M [73]
- Demo G [72]
- Flemming D [70]
- Buschauer R [69]
- Denk T [68]
- Boehringer D [67]
- Xue L [67]
- Loveland AB [66]
- Myasnikov AG [63]
- Fischer P [62]
- Fraser JS [62]
- Lee DJ [61]
- Brown ZP [60]
- Brown A [59]
- Leibundgut M [59]
- Falquet L [57]
- Fromm L [57]
- Hashem Y [57]
- Tegunov D [57]
- Pellegrino J [56]
- Schacherl M [56]
- Sprink T [56]
- Pestova TV [55]
- Seiple IB [55]
- Inada T [54]
- Li X [54]
- Hellen CUT [53]
- Beckert B [52]
- Dabrowski M [52]
- Fedry J [52]
- Gagnon MG [52]
- Khusainov I [52]
- Loerke J [52]
- Ameismeier M [51]
- Matsuo Y [51]
- Baumeister W [50]
- Flugel T [49]
- Grigorieff N [49]
- Chen Y [48]
- Ehrenberg M [47]
- Wang C [47]
- Yusupov M [47]
- Abaeva IS [46]
- Crowe-mcauliffe C [46]
- Hegde RS [46]
- Li W [45]
- Unbehaun A [45]
- Bock LV [44]
- De S [44]
- Fernandez IS [44]
- Fluegel T [44]
- Green R [44]
- Ortega J [44]
- Rappsilber J [44]
- Zhang Y [44]
- Fischer N [43]
- Schroeer B [43]
- Diebolder CA [42]
- Guillen Schlippe Yv [42]
- Hauryliuk V [42]
- Jomaa A [42]
- Lenz S [42]
- Zhang J [42]
- Fucini P [41]
- Kreysing JP [41]
- Welsch S [41]
- Cate JHD [40]
- Carragher B [39]
Author by ORCID
- Beckmann R [194]
- Williamson JR [158]
- Klinge S [145]
- Vanden Broeck A [135]
- Cheng J [126]
- Wilson DN [122]
- Lyumkis D [121]
- Korostelev AA [103]
- Hurt E [95]
- Lau B [95]
- Ramakrishnan V [85]
- Amunts A [84]
- Gao N [84]
- Thoms M [83]
- Mahamid J [82]
- Beck M [73]
- Demo G [72]
- Berninghausen O [70]
- Flemming D [69]
- Denk T [68]
- Cramer P [65]
- Li N [64]
- Pfeffer S [63]
- Frank J [61]
- Burger J [61]
- Mielke T [57]
- Fromm L [57]
- Xue L [57]
- Falquet L [56]
- Boehringer D [53]
- Brown A [52]
- Spahn CMT [52]
- Becker T [50]
- Matsuo Y [50]
- Brown ZP [50]
- Khusainov I [48]
- Forster F [46]
- Loerke J [45]
- Hellen CUT [44]
- Pestova TV [44]
- Rappsilber J [43]
- Rabuck-gibbons JN [43]
- Rodnina MV [43]
- Lenz S [42]
- Kreysing JP [41]
- Welsch S [41]
- Grigorieff N [40]
- Fernandez IS [39]
- Gagnon MG [39]
- Hegde RS [38]
- Yokoyama T [38]
- Zimmermann-kogadeeva M [36]
- Bork P [36]
- Des Georges A [36]
- Diebolder CA [35]
- Guillen Schlippe Yv [35]
- Fraser JS [35]
- Flugel T [35]
- Loveland AB [34]
- Scheerer P [33]
- Shao S [33]
- Cate JHD [32]
- Li N [32]
- Sun M [32]
- Buschauer R [32]
- Blanchard SC [32]
- Faller WJ [31]
- Jomaa A [31]
- Plitzko JM [31]
- Hilal T [31]
- Hauryliuk V [31]
- Ebright RH [30]
- Kaelber JT [30]
- Bhushan S [30]
- Muller CW [30]
- Bridges HR [29]
- Itoh Y [29]
- Hirst J [29]
- Wang C [29]
- Molodtsov V [29]
- Turonova B [28]
- Pellegrino J [28]
- Xing H [28]
- Seiple IB [28]
- Ikeuchi K [28]
- Crowe-mcauliffe C [28]
- Taniguchi R [28]
- Bock LV [28]
- Qin B [27]
- Watson ZL [27]
- Nikolay R [27]
- Vieira-vieira CH [26]
- Tesina P [26]
- Myasnikov AG [26]
- Blaza JN [25]
- Mao Y [25]
- Llacer JL [24]
- Hou YM [24]
- Blaha G [24]
- Valle M [24]
Journal
- NAT COMMUN [581]
- SCIENCE [389]
- NATURE [388]
- MOL CELL [250]
- TO BE PUBLISHED [195]
- NUCLEIC ACIDS RES [191]
- ELIFE [176]
- NAT STRUCT MOL BIOL [171]
- PNAS [161]
- STRUCTURE [134]
- CELL [129]
- EMBO J [121]
- BIORXIV [98]
- NAT METHODS [72]
- EMBO REP [71]
- SCI ADV [51]
- CELL REP [47]
- J MOL BIOL [26]
- J STRUCT BIOL [26]
- PLOS BIOL [25]
- PLOS ONE [25]
- RNA [24]
- SCI REP [24]
- ACTA CRYSTALLOGR D STRUCT BIOL [23]
- NAT CHEM BIOL [23]
- MBIO [19]
- J BIOL CHEM [18]
- BIOMOLECULES [17]
- NAT MICROBIOL [17]
- CELL RES [16]
- PROTEIN CELL [13]
- NAT PLANTS [12]
- INT J MOL SCI [8]
- IUCRJ [8]
- PLANT COMMUN. [7]
- ACS CHEM BIOL [6]
- ANTIOXID REDOX SIGNAL [6]
- J.NEUROSCI. [6]
- PROTEIN SCI [6]
- PROTEOMICS [6]
- ACS CENT SCI [5]
- COMMUN BIOL [5]
- MSPHERE [5]
- ACS PHARMACOL TRANSL SCI [4]
- BIOPOLYM CELL [4]
- CELL DISCOV [4]
- J AM CHEM SOC [4]
- J BACTERIOL [4]
- J CELL BIOL [4]
- J.BIOCHEM. [4]
- NAT CHEM [4]
- PNAS NEXUS [4]
- BIOCHEM J [3]
- FRONT MOL BIOSCI [3]
- ISCIENCE [3]
- RES COMPUT MOL BIOL [3]
- STRUCTURE (LONDON, ENGLAND : 1993) [3]
- ULTRAMICROSCOPY [3]
- BIOCHEMISTRY [2]
- CANCER DISCOV [2]
- CHEM MED CHEM [2]
- MICROBIOL SPECTR [2]
- NAT CELL BIOL [2]
- NAT PROTOC [2]
- PROTEINS [2]
- RNA BIOL [2]
- BIOENGINEERING (BASEL) [1]
- CYSTAL RESEARCH AND TECHNOLOGY [1]
- EMERG MICROBES INFECT [1]
- FEBS LETT [1]
- GENOME BIOL EVOL [1]
- J MED CHEM [1]
- LIFE SCI ALLIANCE [1]
- METHODS [1]
- METHODS IN MOLECULAR BIOLOGY (CLIFTON, N.J.) [1]
- METHODS MOL BIOL [1]
- MICROBIOLOGY (READING, ENGL.) [1]
- MOL CELL PROTEOMICS [1]
- MOLECULES [1]
- NANOMATERIALS (BASEL) [1]
- NUC. ACIDS RES. [1]
- NUCLEIC ACIDS RESEARCH [1]
Software
- RELION [1324]
- CRYOSPARC [491]
- CTFFIND [273]
- GCTF [217]
- SPIDER [176]
- FREALIGN [119]
- WARP [105]
- PYTOM [72]
- CHIMERA [52]
- CISTEM [46]
- MDFF [36]
- IMOD [35]
- COOT [33]
- CTFFIND3, RELION [22]
- EMAN [20]
- SPIDER, APPION [17]
- IMAGIC, CUSTOM, SPIDER [16]
- APPION, SPIDER, EMAN [15]
- O [15]
- XMIPP [14]
- CTFFIND3 [13]
- UCSF CHIMERA [13]
- SPIDER, RELION [12]
- EMAN2 [11]
- GAP [11]
- PHENIX [11]
- RELION, CTFFIND3 [11]
- SIGNATURE, CTFFIND3, SPIDER, SPARX [11]
- SPIDER PACKAGE [11]
- UNBLUR, CTFFIND4, FREALIGN [11]
- CHIMERA, COOT [10]
- IMAGIC [10]
- SPIDER, SPARX [10]
- EMAN2, IMAGIC, FREALIGN, RSAMPLE, CTFFIND3 [9]
- SITUS [9]
- EMCLARITY [8]
- SPIDER, PACKAGE [8]
- PEET [7]
- PYTOM, TOM_TOOLBOX, AV3_TOOLBOX [7]
- TOM TOOLBOX [7]
- XMIPP, SPIDER [7]
- AV3, TOM, PYTOM [6]
- SPHIRE [6]
- STOPGAP [6]
- URO [6]
- CHIMERA, CNS [5]
- CHIMERA, COOT, MDFF [5]
- CHIMERA, REFMAC [5]
- DYNAMO [5]
- IMAGIC, EMAN, FREALIGN, SPIDER, GAP [5]
- IMAGIC-5, SPIDER [5]
- SPIDER, IMAGIC, GAP, CNS, XPLOR [5]
- CHIMERA, EMFIT, COOT [4]
- EM [4]
- NAMD [4]
- RSREF2.0 [4]
- SPIDER, SIGNATURE, MAPPOS [4]
- SPIDER/WEB [4]
- XMIPP, FREALIGN [4]
- ARACHNID, CTFFIND3, RELION [3]
- ARACHNID, RELION, SPIDER [3]
- CHIMERA, COOT, REFMAC [3]
- CHIMERA, MDFF [3]
- CNS [3]
- CRYOLO [3]
- CTFFIND3, RELION1.3 [3]
- IMAGIC 5 [3]
- MANUAL [3]
- MOTIONCORR2 [3]
- SIGNATURE, SPIDER [3]
- SIMPLE [3]
- SPIDER AND GAP [3]
- SPIDER XMIPP [3]
- SPIDER, WELLMAP [3]
- TOM, AV3, PYTOM [3]
- CHIMERA, O, COOT [2]
- CRYODRGN [2]
- CTFFIND3, EMAN2.1, RELION1.3 [2]
- CTFFIND3, RELION, 1.2 [2]
- CTFFIND3, RELION, 1.2, 1.3 [2]
- CTFFIND4 [2]
- EMAN2, CTFFIND4, SPIDER, SPARX [2]
- GROMACS [2]
- IMAGIC-5 SPIDER [2]
- IMAGIC-V [2]
- ML3D [2]
- MOLECULAR DYNAMICS BASED FLEXIBLE FITTING MDFF [2]
- PROGRAM O [2]
- PYMOL, MDFF [2]
- S2S, MODERNA, MODELLER, MDFF [2]
- SITUS, LOCALFIT [2]
- SPARX [2]
- TNT [2]
- UCSF-CHIMERA, MDFF [2]
- URO, REFMAC, NMFF [2]
- AMIRA 3.1, TGS SOFTWARE [1]
- APPION [1]
- AV3 [1]
- BKPR [1]
- CCP4 PACKAGE [1]
Microscope
- FEI TITAN KRIOS [2508]
- FEI POLARA 300 [196]
- FEI TALOS ARCTICA [150]
- TFS KRIOS [146]
- FEI TECNAI F20 [138]
- FEI TECNAI F30 [101]
- FEI/PHILIPS CM200FEG [42]
- FEI TECNAI ARCTICA [26]
- FEI TECNAI 20 [20]
- TFS GLACIOS [20]
- FEI TECNAI SPIRIT [18]
- JEOL 2010F [17]
- FEI TITAN [14]
- FEI TECNAI 12 [10]
- JEOL 2200FS [9]
- JEOL 3200FS [8]
- JEOL CRYO ARM 200 [8]
- JEOL CRYO ARM 300 [8]
- TFS TALOS [6]
- FEI/PHILIPS CM300FEG/T [4]
- JEOL 3100FFC [4]
- JEOL 3200FSC [4]
- FEI/PHILIPS CM200FEG/ST [2]
- JEOL 2200FSC [2]
- JEOL KYOTO-3000SFF [2]
- OTHER [2]
- FEI/PHILIPS CM200FEG/SOPHIE [1]
- FEI/PHILIPS CM200FEG/UT [1]
- FEI/PHILIPS EM420 [1]
- JEOL 1400/HR + YPS FEG [1]
- JEOL 2100F [1]
- JEOL 4000EX [1]
Electron Source
Film or detector model
- GATAN K2 SUMMIT (4K X 4K) [1258]
- GATAN K3 (6K X 4K) [525]
- FEI FALCON II (4K X 4K) [369]
- GATAN K3 BIOQUANTUM (6K X 4K) [323]
- FEI FALCON III (4K X 4K) [272]
- KODAK SO-163 FILM [162]
- GATAN K2 QUANTUM (4K X 4K) [104]
- FEI FALCON IV (4K X 4K) [92]
- TVIPS TEMCAM-F416 (4K X 4K) [50]
- FEI EAGLE (4K X 4K) [37]
- GATAN ULTRASCAN 4000 (4K X 4K) [36]
- GATAN K2 (4K X 4K) [34]
- FEI FALCON I (4K X 4K) [25]
- GENERIC TVIPS (4K X 4K) [23]
- TVIPS TEMCAM-F415 (4K X 4K) [21]
- OTHER [20]
- GATAN K2 BASE (4K X 4K) [17]
- DIRECT ELECTRON DE-64 (8K X 8K) [12]
- DIRECT ELECTRON DE-20 (5K X 3K) [10]
- GATAN MULTISCAN [7]
- GENERIC CCD [5]
- GENERIC GATAN (4K X 4K) [5]
- TVIPS TEMCAM-F816 (8K X 8K) [5]
- DIRECT ELECTRON DE-16 (4K X 4K) [4]
- GATAN K2 IS (4K X 4K) [4]
- DIRECT ELECTRON DE-12 (4K X 3K) [3]
- FEI EAGLE (2K X 2K) [3]
- GENERIC GATAN [3]
- KODAK 4489 FILM [2]
- DECTRIS SINGLA (1K X 1K) [1]
- GATAN ULTRASCAN 1000 (2K X 2K) [1]
- GENERIC FILM [1]
- TFS FALCON 4I (4K X 4K) [1]
EMPIAR Experiment type
EMPIAR Image category